4 June 2018. A biotechnology lab devised techniques for genetically engineered bacteria to produce synthetic proteins that can last longer and are more stable than current protein production methods. Researchers from University of Texas in Austin describe their processes in today’s issue of the journal Nature Biotechnology (paid subscription required).
A team from the lab of biochemistry professor Andrew Ellington at UT-Austin is seeking reliable processes for producing synthetic biologic drugs that have longer shelf and activity lives than current protein-based drugs, such as insulin and monoclonal or highly targeted antibodies. Proteins are chains of amino acids, including cysteine, an amino acid that interacts with other compounds in the body’s cells and quickly breaks down the proteins in the drugs.
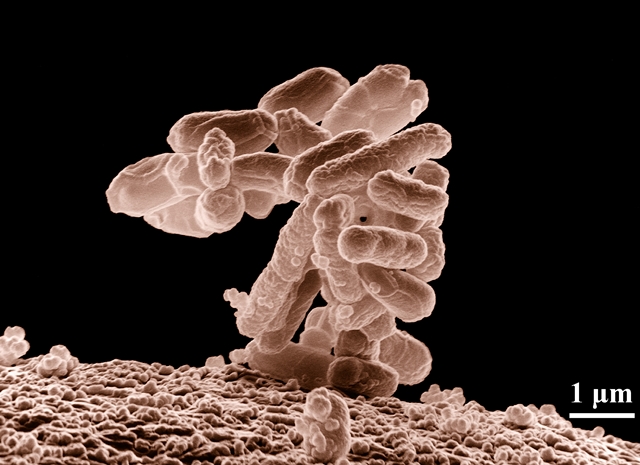
Adding the element selenium to cysteine, however, is known to strengthen and stabilize cysteine while maintaining the amino acid’s functions. The problem, say the researchers, is the lack of a reliable method for producing selenocysteines in sufficient quantities that work well in synthetic proteins. The solution proposed by Ellington and colleagues is to re-engineer the genomes of E. coli bacteria to create a synthetic strain of the bacteria that produces selenocysteines in place of natural cysteine amino acids. While E. coli bacteria are popularly associated with food poisoning, there are many benign strains of the microbe, which are well studied and often used as model organisms in biology labs.
To genetically engineer E. coli, the researchers used a beta-lactamase enzyme often associated with antibiotic-resistant bacteria but with a modified chemistry to bind selenium to cysteine. With this technique the team could create an E. coli strain that produces selenocysteines, but they also directed the evolution of this strain to dominate natural strains in an E. coli community. As a result, say the researchers, they developed a functioning E. coli strain that reliably produces selenocysteine proteins, and in unlimited quantities.
“We have adapted the bacteria’s natural process for inserting selenocysteine to remove all the limitations,” says postdoctoral researcher and first author Ross Thyer in a university statement, “allowing us to recode any position in any protein as a selenocysteine.” The team further engineered E. coli to express fluorescent proteins that illuminate, allowing the researchers to quantify production of selenocysteine proteins. They say the techniques make it possible for the engineered microbes to indefinitely produce selenocysteine proteins in industrial-scale quantities.
Ellington and Thyer are co-founders with Harvard University geneticist and serial entrepreneur George Church and others of the start-up enterprise GRO Biosciences in Cambridge, Massachusetts to produce synthetic proteins from genetically-recoded bacteria, and serve as scientific advisers to the company. In September 2017, GRO Biosciences secured seed capital of $2.1 million and announced plans to produce therapeutic proteins with expanded potency and stability. As reported by Science & Enterprise in August 2016, Church and colleagues from another company he founded pioneered techniques for streamlining the E. coli genome, removing redundant elements.
More from Science & Enterprise:
- Trial Testing Gene-Edited, Modified Oilseed Plants
- Genetic Repair Company Starts-Up, Gains $87 Million
- Cellectis, Wyss Inst. Partner on Recoding Human Genome
- Platform to Integrate Robotics, Synthetic Biology, A.I.
- Cough Suppressant Produced from Engineered Yeast Cells
* * *

 RSS - Posts
RSS - Posts
[…] Bacterial Genome Recoded to Produce Stable Protein Drugs […]